What specific regions or genes does the StrainSEEK® 3Mb panel cover?
The cannabis nomenclature for gene names is not well defined, but we provide a comprehensive coverage analysis of our StrainSEEK® 3 Mb panel here.
Cannabis nomenclature for gene names is not well defined. Due to its long history of prohibition, scientists have not been able to study the cannabis genome in depth until recently. Now that research is exploding and with it our understanding of how beautifully complex Cannabis is.
Our analysis includes 182 different cannabis samples which were sequenced with our StrainSEEK® 3Mb (actually 3.2Mb) panel. These 182 samples have all been deposited into Kannapedia and are part of our publicly accessible StrainSEEK® database.
Coverage was calculated at every position of our JLion_V6_7M_polarstar_purged.fasta reference sequence that has NGS coverage in our panel for all 182 samples (NOTE: this reference sequence along with the gene annotation was deposited into NCBI and is publicly available here: https://www.ncbi.nlm.nih.gov/nuccore/JAATIP000000000). Next, we took our list of gene annotations that are mapped to this reference sequence (27,664 genes) and determined the following:
The percentage of each region where 1% and 10% of the samples are collected for C1, C10, C20, C30 and C50 (so there are 10 coverage columns added to the spreadsheet). C1 are the total bases with >=1x sequencing coverage, C10 is the total bases over >=10x sequencing coverage, and so on up to C50. To illustrate how this table should be interpreted, let's look at the first 2 rows of the table (out of the the total 27,664):
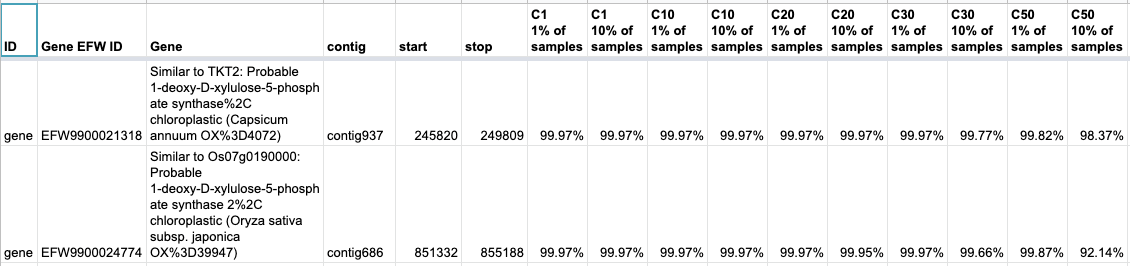
The first gene with the unique ID EFW9900021318 maps to positions 245,820-249,809 on contig937 of our JLion_V6_7M_polarstar_purged.fasta reference sequence. This is a 3,990bp region. 10% or more of the samples (so, >=19 of the 182 samples) have >=50x coverage at 98.37% of this 4kb region (so ~3.92kb). That is the value in ColumnP of our gene coverage worksheet.
If you are interested in obtaining the complete list of 27,664 genes covered by 182 samples of our StrainSEEK® 3Mb panel please complete our contact form.
